PRIMED: PRIMEr Database for Deleting and Tagging All Fission and Budding Yeast Genes Developed Using the Open-Source Genome Retrieval Script (GRS)
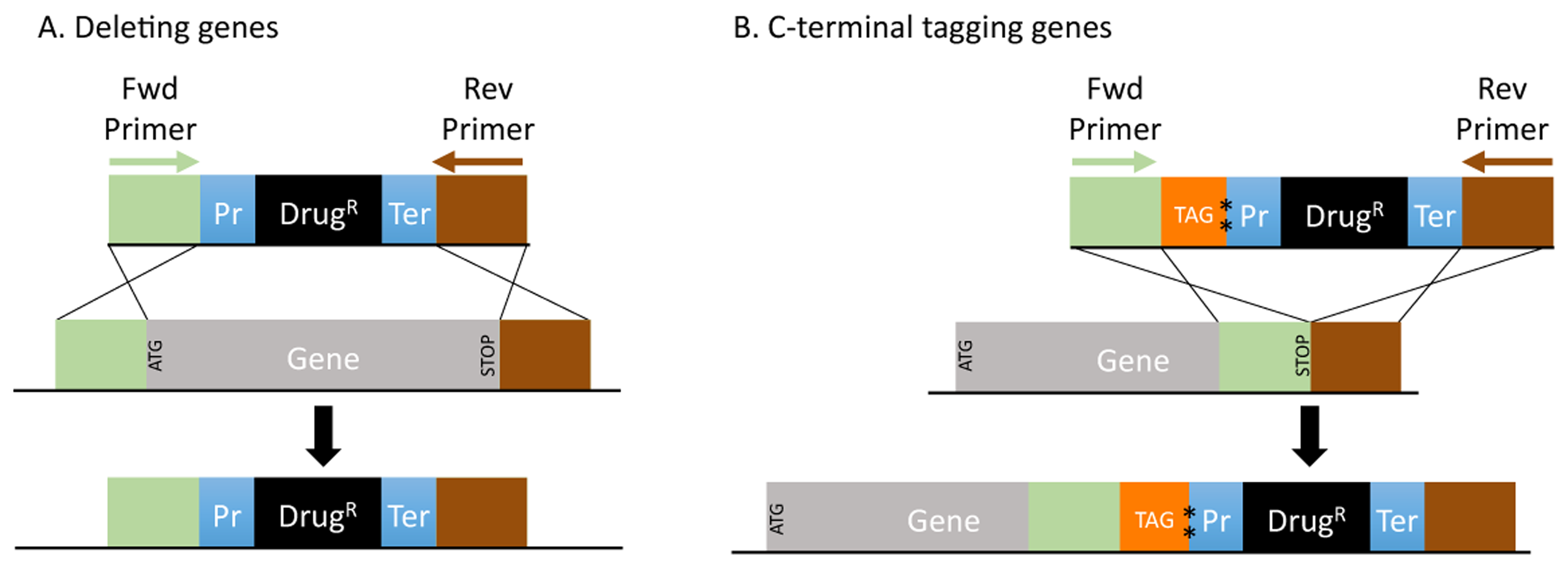
The fission (Schizosaccharomyces pombe) and budding (Saccharomyces cerevisiae) yeasts have served as excellent models for many seminal discoveries in eukaryotic biology. In these organisms, genes are deleted or tagged easily by transforming cells with PCR-generated DNA inserts, flanked by short (50-100bp) regions of gene homology. These PCR reactions use especially designed long primers, which, in addition to the priming sites, carry homology for gene targeting. Primer design follows a fixed method but is tedious and time-consuming especially when done for a large number of genes. To automate this process, we developed the Python-based Genome Retrieval Script (GRS), an easily customizable open-source script for genome analysis. Using GRS, we created PRIMED, the complete PRIMEr D atabase for deleting and C-terminal tagging genes in the main S. pombe and five of the most commonly used S. cerevisiae strains. Because of the importance of noncoding RNAs (ncRNAs) in many biological processes, we also included the deletion primer set for these features in each genome. PRIMED are accurate and comprehensive and are provided as downloadable Excel files, removing the need for future primer design, especially for large-scale functional analyses. Furthermore, the open-source GRS can be used broadly to retrieve genome information from custom or other annotated genomes, thus providing a suitable platform for building other genomic tools by the yeast or other research communities.